Kyle's BIMM 143 Portfolio

My classwork for BIMM143
Class 12 Pt. 2
Kyle Canturia (A17502778)
Section 1: Identify genetic variants of interest
Downloaded data from Ensembl as a csv file
mxl <- read.csv("373531-SampleGenotypes-Homo_sapiens_Variation_Sample_rs8067378.csv")
head(mxl)
Sample..Male.Female.Unknown. Genotype..forward.strand. Population.s. Father
1 NA19648 (F) A|A ALL, AMR, MXL -
2 NA19649 (M) G|G ALL, AMR, MXL -
3 NA19651 (F) A|A ALL, AMR, MXL -
4 NA19652 (M) G|G ALL, AMR, MXL -
5 NA19654 (F) G|G ALL, AMR, MXL -
6 NA19655 (M) A|G ALL, AMR, MXL -
Mother
1 -
2 -
3 -
4 -
5 -
6 -
table(mxl$Genotype..forward.strand.)
A|A A|G G|A G|G
22 21 12 9
GBR dataset
gbr <- read.csv("373522-SampleGenotypes-Homo_sapiens_Variation_Sample_rs8067378.csv")
head(gbr)
Sample..Male.Female.Unknown. Genotype..forward.strand. Population.s. Father
1 HG00096 (M) A|A ALL, EUR, GBR -
2 HG00097 (F) G|A ALL, EUR, GBR -
3 HG00099 (F) G|G ALL, EUR, GBR -
4 HG00100 (F) A|A ALL, EUR, GBR -
5 HG00101 (M) A|A ALL, EUR, GBR -
6 HG00102 (F) A|A ALL, EUR, GBR -
Mother
1 -
2 -
3 -
4 -
5 -
6 -
Section 4: Population Scale Analysis
pop <- read.table("rs8067378_ENSG00000172057.6.txt")
head(pop)
sample geno exp
1 HG00367 A/G 28.96038
2 NA20768 A/G 20.24449
3 HG00361 A/A 31.32628
4 HG00135 A/A 34.11169
5 NA18870 G/G 18.25141
6 NA11993 A/A 32.89721
#reads dataset and assigns it to a variable
Q13: Read this file into R and determine the sample size for each genotype and their corresponding median expression levels for each of these genotypes.
table(pop$geno)
A/A A/G G/G
108 233 121
##only looks at the column "geno" and returns count of how many of each genotype is present
Theres 108 A|A, 233 A|G, and 121 G|G.
library(dplyr)
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
pop %>%
group_by(geno) %>%
summarize(geno_median = median(exp))
# A tibble: 3 × 2
geno geno_median
<chr> <dbl>
1 A/A 31.2
2 A/G 25.1
3 G/G 20.1
##uses dplyr tools to first group the samples by genotype, then summarizes the median of each genotype
The median expression levels for A|A, A|G, and G|G are 31.24, 25.06, and 20.07 respectively.
Q14: Generate a boxplot with a box per genotype, what could you infer from the relative expression value between A/A and G/G displayed in this plot? Does the SNP effect the expression of ORMDL3?
library(ggplot2)
ggplot(pop) +
aes(geno, exp) +
geom_boxplot()
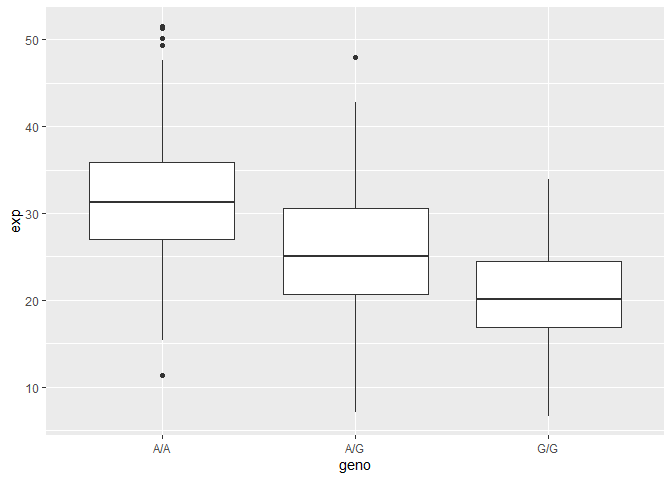
##makes boxplot with geno on the x axis and exp on the y
A|A has the highest expression value while G|G has the lowest, and A|G is in between. From the plot, it’s evident that the SNP is associated with differing expressions of ORMDL3.